Tribe is a collaborative platform that allows mining of big data across bioinformatics systems. Here at the NDEx Project, we are excited to find a system with a similar philosophy and complementary content. Tribe has features that make it an interesting resource for both biology and bioinformatics researchers and we look forward to evaluating it in greater depth.
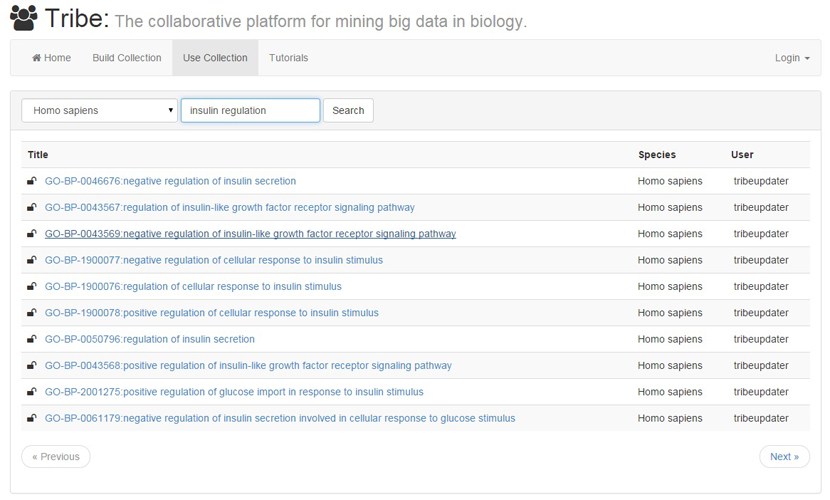
Centered on the idea of collaborative data mining, Tribe allows researchers to collaborate, store their findings in one place and use these collections to mine large scale data sources.
Tribe has a built-in permission system that allows to control the level of privacy of each collection and grant access to specific users for collaboration.
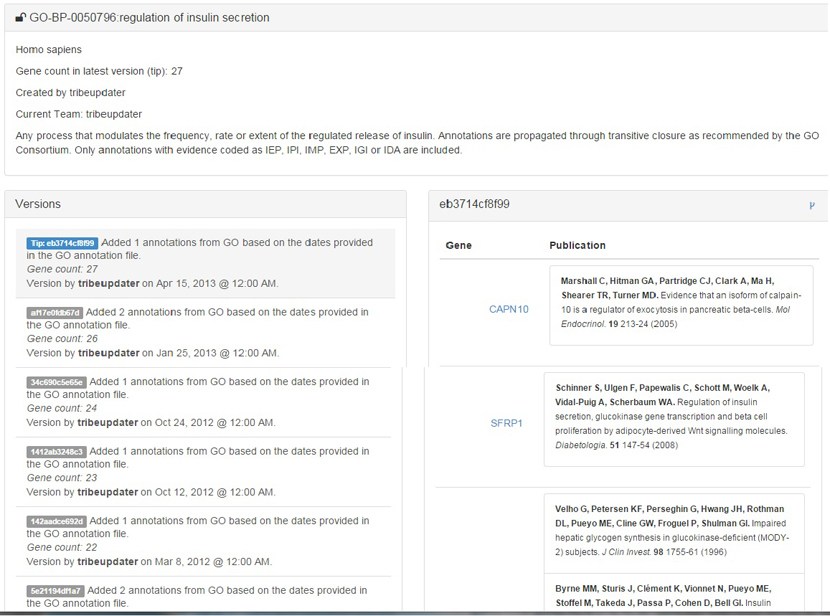
A versioning system (picture below) ensures that all information regarding collections are stored and always accessible: a forking feature, similar to the one used by popular code repositories such as GitHub, allows to branch off and create a new knowledgebase to add your own findings and share with collaborators.
The Tribe API returns JSON objects thus allowing you to use your favorite scripting language; in addition, the Tribe server handles gene name mapping, so all collections can be returned with gene identifiers that you need. Tribe also supports the OAuth2 protocol making it easy to link to any webserver: this way, users can access their Tribe collections right from their own servers.
Tribe was developed by the Greene Laboratory at the Geisel School of Medicine, Dartmouth College.