CyNDEx – The NDEx Cytoscape App – Tutorial
Last updated: February 1st, 2018
Overview
The NDEx Cytoscape App (CyNDEx) provides a mechanism for you to take any network in Cytoscape and upload it to NDEx. You can also download networks from NDEx to Cytoscape. The goal of CyNDEx is to be both easy to install and easy to use. If you have any ideas for how it could be even better, we want to hear from you!
CyNDEx was developed by the NDEx Project at UC San Diego and is available for download on the Cytoscape App Store.
Why Use This App?
NDEx provides a great location on the Internet where you can both save, share and publish your networks! You can think of it as a sort of Google Drive for your networks, only with a lot more network-centric power. CyNDEx allows you to transfer your networks between Cytoscape, the leading desktop solution for visualizing biological networks, and NDEx. Although we have biological networks in mind, any kind of network can be stored in NDEx.
Getting Started
Before you can install CyNDEx, you must have Cytoscape 3.3.0 or later installed along with 64-bit Java 8. You also must ensure that your JAVA_HOME environmental variable is set to Java 8.
- To install Java 8 or later, go here.
- For instruction on how to set JAVA_HOME, go here.
- To install Cytoscape 3.3.0 or later, go here.
Installing CyNDEx
Now that you have Cytoscape installed and Java 8 setup correctly, it is time to install CyNDEx. The CyNDEx App can be downloaded from the Cytoscape App store. Open up Cytoscape and select App Manager from the Apps menu…
Choose the Install Apps tab, select CyNDEx from the list of available apps and click the Install button. Alternatively, if you already had CyNDEx installed, you can select the Check for Updates tab and install the latest version available.

Once the App Manager will indicate that the status of CyNDEx is installed, you can close the App Manager.
Now, under the Apps menu in Cytoscape, you will see an NDEx menu with 3 options.
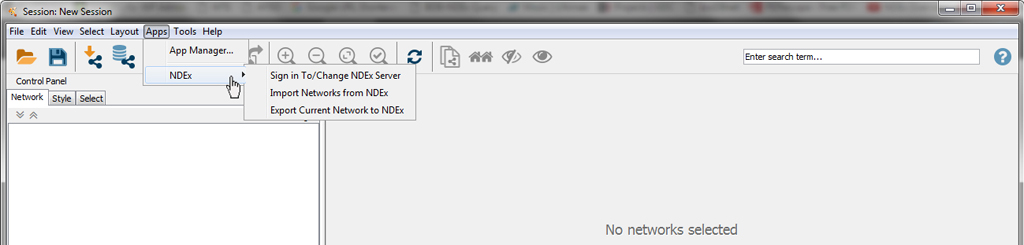
Using CyNDEx
By default, you can anonymously download any "public" networks from the PUBLIC NDEx SERVER to Cytoscape using CyNDEx you just installed; however, you will not be able to download "private" networks or upload any networks from Cytoscape to NDEx unless you are logged in to your NDEx Account. Therefore, in order to use the full functionalities of CyNDEx, you need an NDEx account. This tutorial was designed assuming that you already have an NDEx Account.
For instructions on how to create and NDEx Account, go here. Once you have registered your NDEx Account, proceed with the following instructons.
– Log in to NDEx
In Cytoscape, click the Apps menu, select NDEx and then click Sign In To/Change NDEx Server…
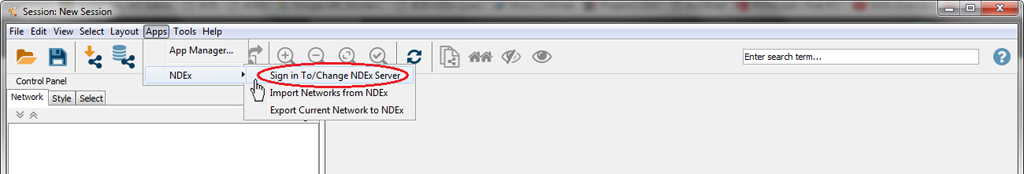
Make sure that NDEx Public is selected and click Edit

Enter your username and password and click Save

Finally, click the Connect button in the bottom right corner and you will get a confirmation message that indicates that your username and password was correctly recognized.
– Download (import) networks from NDEx
Choose the Import Networks from NDEx menu item.
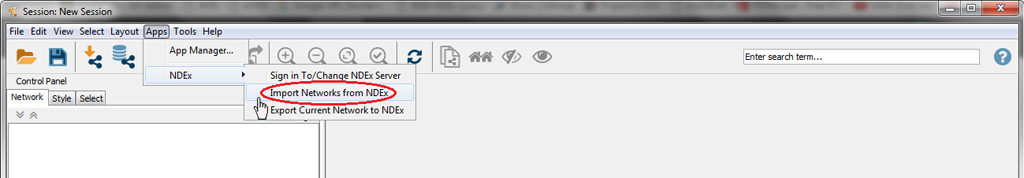
On the Find Networks screen, you can either select one of the networks on the list or search for more specific networks. For example, you can type the word "metabolism" in the text box and click Search. Then make sure the "Metabolism of RNA" network is selected and press the Load Network button.

The network will begin to download, but the dialog will remain open so that you can select addition networks to download into Cytoscape. You may only download one network at a time, but it is sometimes possible to get the next network started while you are waiting for the previous one to finish.
We are done loading networks and now want to view them in Cytoscape. Click the Done Loading Networks button in the bottom right corner so that we can see what we have imported in Cytoscape.

– Visualize downloaded networks in Cytoscape (optional)
In some cases, a downloaded network could look like it consists of 1 single node… The reason for this odd behavior is that the downloaded network doesn't have any presentation properties set and therefore it appears in a simplified manner in Cytoscape. In order to properly visualize such a network, we need to apply a visualization layout. Note: this step is only necessary when importing networks larger than 10,000 edges; for networks smaller than 10,000 edges, a default layout is automatically applied when the network is loaded in Cytoscape.
To apply a layout, just select your network, open the Cytoscape Layout menu and choose your favorite layout item. In this example we apply the Prefuse Force Directed Layout wich is also the default layout. As our imported network is smaller, the default layout has already been applied and this step will have no effect.

Upload (export) networks to NDEx
Once you are done working on your network in Cytoscape, it is time to perform the final step in our tutorial: exporting a network to NDEx.
Although we illustrate how to upload the "Metabolism of RNA" network, the same exact procedure can be applied to any other networks you have been working on in Cytoscape, regardless of their provenance. Let's get started!
First, select the "Metabolism of RNA" network by clicking on its title bar, then open the Apps menu, select NDEx and click on Export Current Network to NDEx.
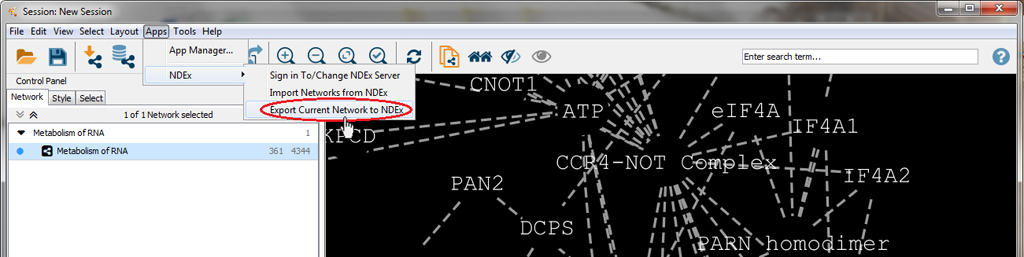
A window will appear which tells you which NDEx SERVER you are currently connected to, what users account that network will be exported to as well as the size of the network.
Here, you can change the network name as well as decide whether you want to upload a new copy of the network (default behavior) or update the existing one using the checkbox indicated by the red arrow. Please note that in order to update a network, you must have either "Edit" or "Admin" permission on that network. For additional information about network permissions in NDEx please review the guide about Obtaining and Granting Access to a Network.
In this tutorial, we will leave the name of the network unchanged and click the "Upload Network to NDEx" button thus creating a fresh copy of the network. Note: during the upload procedure the dialog window will not show any progress bar; uploading large networks will require some time, and the export window will close automatically once the export process has completed.

Done! You can now log in to your NDEx account and see that your "Metabolism of RNA" network is displayed in your personal account page.
With NDEx and CyNDEx, you can easily share your networks with collaborators as well as use and improve networks shared by others. Tutorials and detailed user documentation for NDEx is available in the Documentation page on our website.
We hope you enjoyed this tutorial and the power you now have at your fingertips.